Research
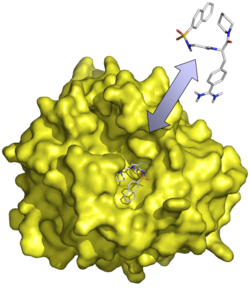
General topic of our research is the investigation of protein-ligand interactions by means of computer-based methods. Our work may be divided in application projects on the one hand and methods development on the other.
In application projects, methods such as docking, scoring and molecular dynamics (MD) simulations are used to analyse protein-ligand complexes, to support the interpretation of experimental results, and to establish hypotheses regarding structural and energetic aspects of the interactions. In cases where finding new ligands is the immediate goal, various approaches for virtual screening of molecular databases are used. A large part of this work serves efforts of ligand design for protein targets of pharmaceutical interest and is, thus, frequently embedded in cooperations with experimental groups.
Projects of method development are intended to improve and advance existing computational methods in drug design. Although these methods have reached an impressive performance over the last years, a series of problems still persist. For example, the prediction of affinity and other thermodynamic quantities for biomolecular complexes can often not yet be made with sufficient speed and reliability. The conformational flexibility of proteins may well be investigated by time-consuming MD simulations, but consideration of this flexibility in docking, scoring, and virtual screening is still not straightforward. Finally, also the adequate treatment of water and solvation effects in the context of protein-ligand interactions causes major problems. Accordingly, methodically oriented projects aim to make contributions to solve these problems.